2016-05-30 Scaling up a scientific computation
This article was written with Aaron Contorer and originally appeared as an experience report on the FPComplete website.
Abstract
Often, complex scientific computations are created on a researcher’s desktop computer, then dramatically refactored – partly or completely rewritten – to run much faster on large servers, clusters, or clouds. Moving a computation to a another computer is easy, but changing it to run faster and faster (to support bigger and bigger datasets) can be hard. In heavily regulated industries the cost of refactors is often very high, given the high cost of changing code. The goal is to achieve great throughput while showing that the computation is still correct.
A client of FP Complete faced this issue while improving the performance of a software-based medical device. We dramatically improved its speed and scalability by building a framework to distribute the workload efficiently across multiple processes on multiple CPU cores, and then across multiple machines. We included the “middleware” infrastructure supporting these improvements as part of FP Complete’s High Performance Computing framework, allowing for easy deployment of the scaled-up device on Amazon’s AWS cloud computing platform.
Client Situation and Need
The Haskell programming language, designed for reliability, productivity, and scalability, is one of very few languages to be purely functional. Code with side effects is explicitly separated from the “pure” functions that always return the same output give the same input. This makes Haskell very appealing to people in need of implementing and verifying a mathematical specification, since pure functions are much easier to handle formally. After all, mathematics is only concerned with pure functions.
Our client chose Haskell to implement a new software-based medical device, processing patient data and performing unusually detailed predictive modeling to improve the treatment of patients with a targeted disease. The whole core technology can be seen as a single pure function – patient data in, predictive analysis out. While the client used the power of Haskell to get the software working – they had a safe and effective prototype device – the vast amount of computation involved meant that it would need multiple CPU cores to run quickly. And in theory, the use of Haskell should have meant they could just “turn on” native multicore support for more throughput.
But despite their use of Haskell’s native parallel/multicore programming features, the performance was not linearly improving as more CPUs were added. We set to investigate why the software was not scaling on a single machine as they had hoped, and if possible allow scaling to much larger (18 core) machines and even beyond a single machine to a cloud-based “compute cluster” of big servers.
Overview of the solution
FP Complete worked around this problem by developing a platform capable of easily distributing Haskell computation across distinct processes. This allowed us not only to improve performance by 100% on a single 18-core machine, but also to distribute the computation across multiple machines, achieving speedups of up to 600% compared to the best performance we could get with the single-process version.
We choose this approach since it allowed us to achieve a dramatic speedup without any change to the core algorithm, and maintaining the output of the device byte-identical to the original version at all times, thus avoiding the burden of verifying the output twice, given the highly regulated environment the device operates in.
Detailed Analysis
We used AWS to create virtual machines with differing numbers of CPU cores, and profiled the application’s behavior on each.
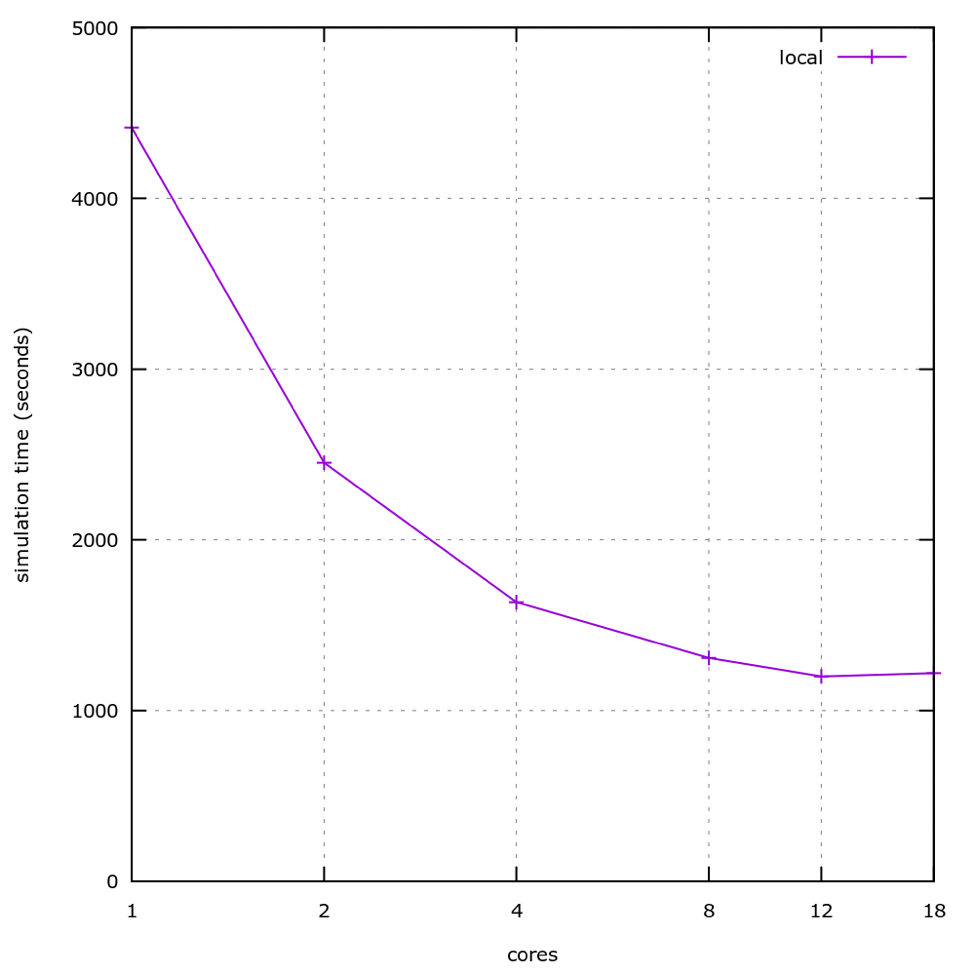
As seen here, performance stops improving significantly past 4 cores, and in fact stops improving at all approaching 10 cores. Further investigation actually showed slower performance above 12 cores.
As a memory-managed language, Haskell relies critically on garbage collection – the recycling of unused memory space. Knowing the importance of fast garbage collection to good performance, and knowing that our client’s application used and released large amounts of memory, we suspected the problem might be here.
We investigated further and found inefficient behavior of the Haskell garbage collector when managing a single heap accessed by many concurrent threads. To establish the source of the problem we took advantage of the instrumentation in the Glasgow Haskell Compiler runtime. Specifically, we were able to precisely measure how much time was spent garbage collecting, and how parallelized the garbage collector itself was. We discovered that as we added CPUs garbage collection took a higher and higher percentage of the total time – up to 60% of the times in some cases, compared to less than 10% with one CPU. Using a tool called ThreadScope, it became clear that as the number of CPUs grew, the garbage collection pauses would become more and more frequent (since memory would be allocated faster, given the parallel work). The pauses would also be longer, given that they had to traverse a larger heap. This, coupled with each garbage collection pause blocking all the threads each time, caused the slowdown our client was experiencing.
Usually these problems are solvable using data structures that exert less pressure on the garbage collector. However, this kind of restructuring would have required broad changes to how the core algorithm of the device are structured, incurring significant costs and delays, and possibly complicating the regulatory filing. We needed to find a solution to the scalability problem without modifying the client’s core code.
Solution details
Since the inefficiencies we were experiencing were due to Haskell’s garbage collector having trouble managing a very big heap accessed by multiple threads, one way to mitigate the problem is to split up the work in many small heaps serving different threads or processes. Thus, garbage collection in one memory heap would not block users of the other heaps. We decided to distribute the workload across several separate single-threaded processes, each with its own heap, so that each would be very productive while still being able to distribute work across vast computing resources.
To distribute work efficiently, we developed a design using a master-slave architecture with the master orchestrating work to slaves. The slaves communicate with the master via messages over TCP sockets, and thus can be on the same machine or on separate machines – a “two for one” win since the same code could support multi-core as well as multi-machine scaling. We distributed large chunks of work per message, to ensure that the latency in message-passing would not significantly affect performance.
The client’s predictive model uses a statistical approach called time-series Monte Carlo, which explores thousands of possible evolutions over time. Thus as a first attempt the master would orchestrate the work as follows:
The master node is initialized with the initial states
When the master node needs to evolve a state, it transfers it to a slave node, which can be another process on the same machine, or on another machine across a network connection
The slave node executes the compute-intensive “evolution” function, and then returns the new state to the master
The master inspects the new states as they come from the slaves and continues evolving them
The problem with this first approach is that the states are relatively big – in the hundreds of kilobytes per state, for a total of several gigabytes of states alive at any given time. This makes transferring them to a slave each time impractical.
We developed an algorithm capable of persisting the states on slave nodes, and then evolving them directly on the slaves without having to transfer them each time. Moreover, the master also take care to dynamically rebalance the work when a slave ends up having a backlog of states to evolve while others are idling. This keeps the communication traffic to a minimum while achieving good work balance across slave nodes. Crucially, all this management was handled outside the core regulated code, the mathematical analysis.
A remaining bottleneck was converting Haskell objects to streams of bytes that could be transferred as a message. This led to optimization work that culminated in the development of a new serialization library called “store” which improves the Haskell status quo by several orders of magnitude. With the client’s permission we released this library under a permissive open source license, following a string of FP Complete releases aiming to improve the Haskell ecosystem while protecting client confidentiality. This helps to reduce future maintenance work for the client, by bringing the open-source community to bear on maintaining the generic function over time.
Results and Benefits
At the end of this optimization effort we doubled the performance of the device on the 18-core machines that we were using, and enabled us to gain a speedup of up to 500% by distributing the work across several machines, after which we hit other limits that we are still improving on.
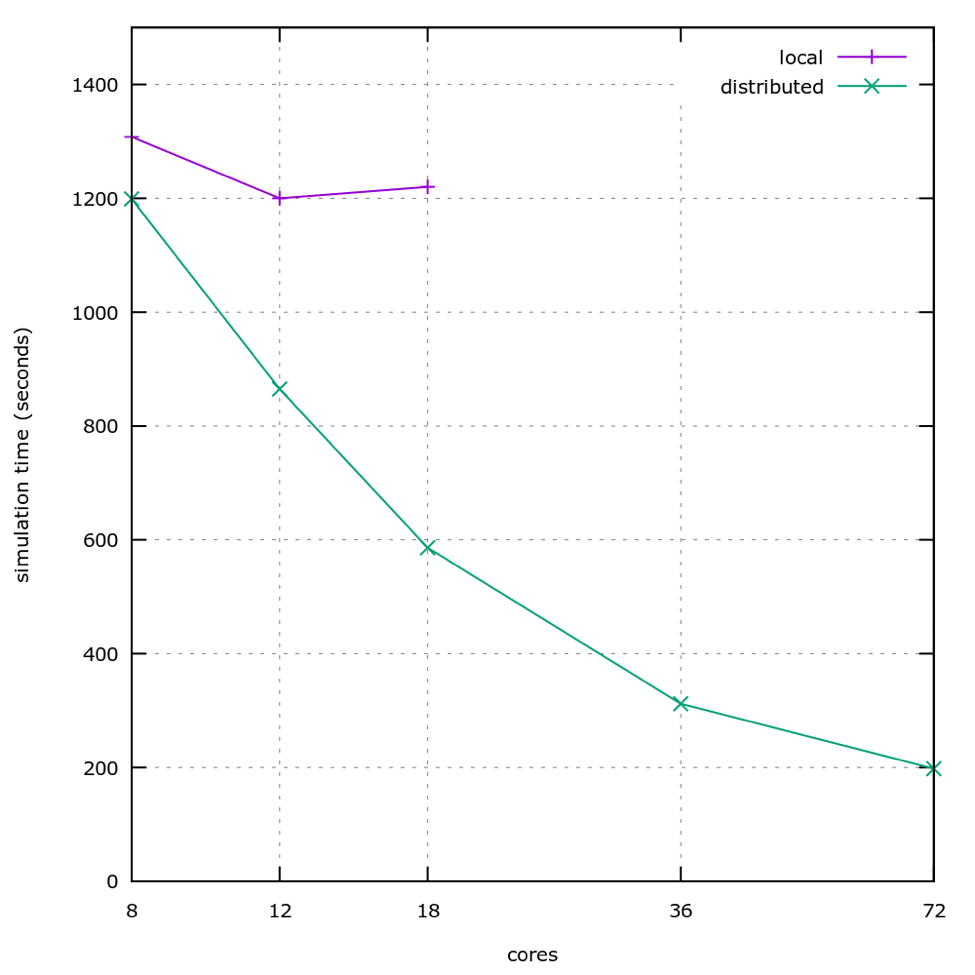
As the graph above shows, the distributed version of the software is already twice as fast on a single, 18 cores machine, and is then capable of scaling beyond one machine for even faster simulations. For example, the plot shows the time the simulation takes for 2 machines (36 cores) and 4 machines (72 cores), where the simulation time is slightly more than 3 minutes, while the best time we could get using the local version of the software is around 20 minutes. This is not only advantageous for the product, but also a boon to development: programmers testing changes or data scientists tuning parameters can iterate 6 times faster.
Moreover, we integrated this platform to distribute work into a broader high-performance computing (HPC) framework that allows the client to easily and reliably schedule requests and wait for responses. This framework allows multiple clients, even across a remote Web API, to schedule work that is automatically and flexibly distributed across a cluster. The cluster can expand and shrink with no downtime, creating and deleting cloud compute machines to quickly react to changing demands. This HPC framework is architected to be extremely easily to deploy and scale, taking advantage of the AWS computing platform (both in the form of the EC2 and ElastiCache) and Docker containers to maximize reliability and minimize maintenance costs.
Having a direct handle on how the work is distributed let us integrate the strategy to distribute work into the client’s broader strategy to deploy the medical device reliably and scalably. This means that we’re able to use the hardware we are paying for as efficiently as possible, and we’re able to scale seamlessly as the load increases.